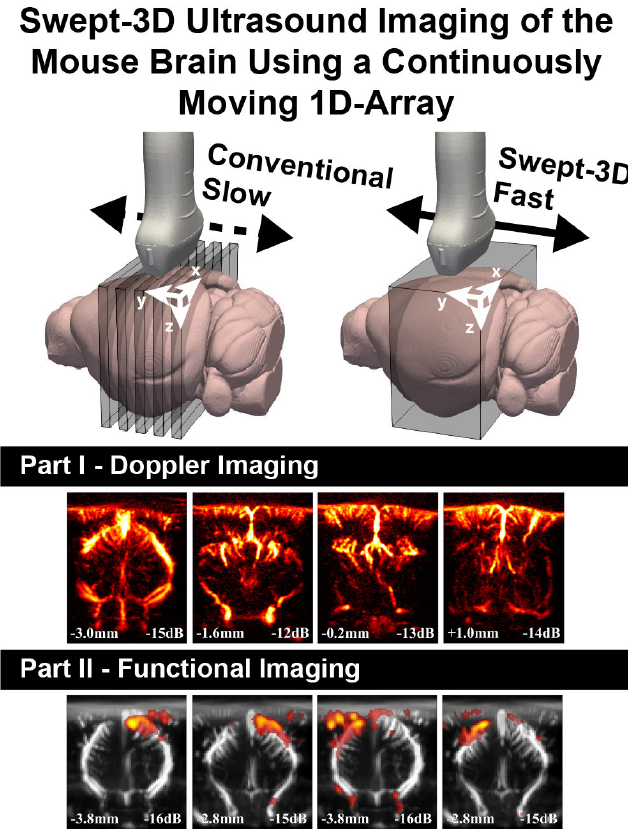
Swept-3-D Ultrasound Imaging of the Mouse Brain Using a Continuously Moving 1-D-Array—Part I: Doppler Imaging
Volumetric 3-D Doppler ultrasound imaging can be used to investigate large scale blood dynamics outside of the limited view that conventional 2-D power Doppler images (PDIs) provide. To create 3-D PDIs, 2-D-matrix array transducers can be used to insonify a large volume for every transmission; however, these matrices suffer from low sensitivity, high complexity, and high cost. More typically, a 1-D-array transducer is used to scan a series of stationary 2-D PDIs, after which a 3-D volume is created by concatenating the 2-D PDIs in postprocessing, which results in long scan times due to repeated measurements. Our objective was to achieve volumetric 3-D Doppler ultrasound imaging with a high Doppler sensitivity, similar to that of a typical stationary recording using a 1-D-array transducer, while being more affordable than using 2-D-matrix arrays. We achieved this by mounting a 1-D-array transducer to a high-precision motorized linear stage and continuously translating over the mouse brain in a sweeping manner. For Part I of this article, we focused on creating the best vascular images by investigating how to best combine filtered beamformed ultrasound frames, which were not acquired at the same spatial locations, into PDIs. Part II focuses on the implications of sampling transient brain hemodynamics through functional ultrasound (fUS) while continuously translating over the mouse brain. In Part I, we show how the speed at which we sweep our 1-D-array transducer affects the Doppler spectrum in a flow phantom. In vivo recordings were performed on the mouse brain while varying the sweeping speed, showing how higher sweeping speeds negatively affect the PDI quality. A weighting vector is found to combine frames while continuously moving over the mouse brain, allowing us to create swept PDIs of similar sensitivity when compared with those obtained using a stationary 1-D-array while allowing a significantly higher 3-D Doppler volume rate and maintaining the benefits of having a low computational and monetary cost. We show that a vascular subvolume of 6 mm can be scanned in 2.5 s, with a PDI reconstructed every $200 \mu \text{m}$ , outperforming classical staged recording methods.
Chris de Zeeuw
10.1109/TUFFC.2023.3318653
3-D Doppler imaging, 3-D mouse brain, Doppler flow phantom, motorized linear stage, volumetric Doppler imaging